Something I’ve learned this summer is that things always take longer than expected (at least in the case of this project). For the past 8 or so weeks, I’ve been working designing primers that will successfully amplify the genes we want to study, something that I had originally planned to finish by week 4 at latest. Though initially this was pretty stressful and a little frustrating, it’s given me the opportunity to troubleshoot by changing steps of the procedure a little at a time to pinpoint what went wrong.
If you’re interested in more details on what I’ve learned about primer design, read on…
How do primers work?
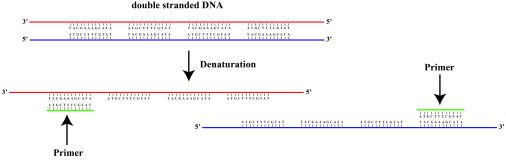
Primers are used in PCR, a process that amplifies a certain gene. During PCR, the two strands that make up DNA are melted so that they come apart. Primers are short strands of nucleotides that bind to a single strand of DNA. Both a forward and reverse primer are used, one binding to each of the two DNA strands, so that the region you want to amplify is bounded by primers. This allows an enzyme (Taq Polymerase) to lengthen the primer into a full strand, creating a complete, double-stranded segment of DNA, which is your PCR product.
The difficulty with Carbonic Anhydrase primers:
Our project aims to amplify genes coding for Carbonic Anhydrase (CA) enzymes. However, CA genes are found in many different organisms (from bacteria to humans!), and even though they all code for the same type of protein, the proteins and the genes themselves are very diverse. So if we could look at all the CA from all the different microorganisms in soil (which is what we want to do, but can’t as of now), the genes would all look very different. This makes it difficult (actually, probably impossible) to find a region that’s common among all CA genes where the primers can bind to.
The solution:
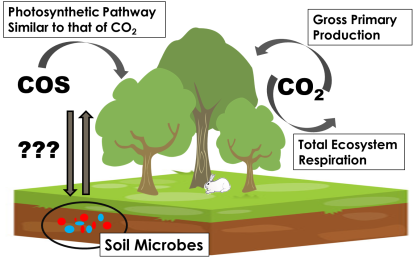
We ended up designing primers for a very specific type of CA, called Cosase by the researchers who discovered it (Ogawa et al., 2013). They predicted that Cosase was responsible for degrading of COS, so we want to see if there’s enough Cosase in our soils for it to feasibly be responsible for all or most of the COS uptake exhibited by our soil samples. There are relatively few Cosase genes in the NCBI database of sequenced genomes (the organisms whose genetic makeup is completely known), so it will be interesting if we find new, previously unknown Cosase genes!